FDIF (Formula-Driven supervised learning with Implicit Functions) is a novel pre-training framework for 3D medical image segmentation that generates synthetic training data entirely from mathematical formulas. By utilizing Signed Distance Functions (SDFs), it achieves SOTA performance among formula-driven methods and matches the results of self-supervised learning (SSL) models trained on massive real-world datasets.
TL;DR
Researchers have developed FDIF (Formula-Driven supervised learning with Implicit Functions), a framework that allows AI models to learn 3D medical image segmentation without ever seeing a real human organ during pre-training. By using Signed Distance Functions (SDFs) to procedurally generate complex shapes and textures, FDIF matches or beats SOTA self-supervised methods that rely on over 100,000 real medical volumes.
Background: The Privacy-Performance Paradox
In the world of 3D medical imaging (CT/MRI), we face a "Goldilocks" problem: AI needs massive datasets to perform well, but privacy laws (like HIPAA/GDPR) and the grueling cost of expert annotation make those datasets nearly impossible to share at scale.
While Self-Supervised Learning (SSL) has been the go-to solution, it still requires "unlabeled" real data, which is still subject to privacy constraints. Formula-Driven Supervised Learning (FDSL) offers a radical exit: Generate everything—images and labels—from pure math.
The Core Innovation: Why Implicit Functions?
Previous FDSL attempts like PrimGeoSeg were "voxel-based," essentially stacking 2D slices. This limited them to simple extruded shapes. FDIF shifts the paradigm to Implicit Functions, specifically Signed Distance Functions (SDFs).
1. Geometric Expressiveness
Unlike voxels, SDFs define a surface as the zero level-set of a continuous mathematical function. This allows for:
- Complex Topologies: Easily representing holes, cavities, and organic intersections.
- Infinite Resolution: No more "blocky" artifacts during the generation phase.
2. The Texture Breakthrough: DF & MF
Medical segmentation isn't just about shape; it's about the "look" of the tissue. FDIF introduces:
- Displacement Functions (DF): These add "noise" or "ridges" to the surface of the mathematical shape, simulating the rough edges of tumors or the branching of vessels.
- Mapper Functions (MF): These map the distance from the boundary to specific intensities. This allows the model to learn "intensity gradients"—crucial for identifying where one organ ends and another begins.
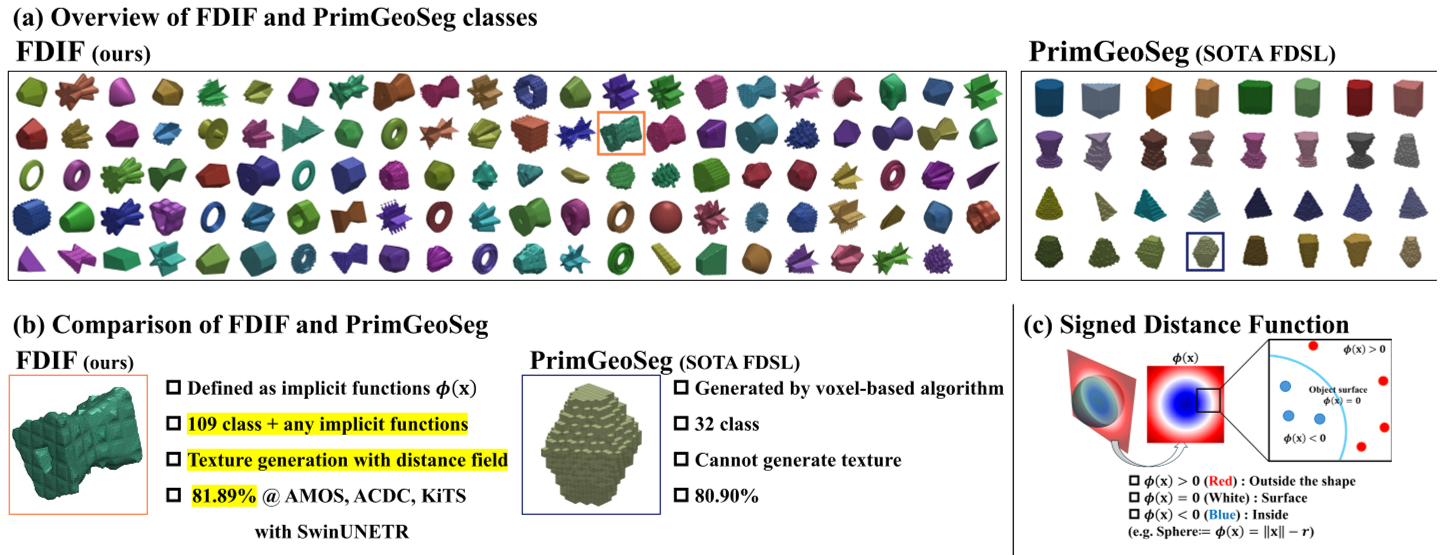 Figure 1: Comparison between previous voxel-based generation (a, b) and the proposed SDF-based FDIF (c), which supports complex topologies and surface-aware textures.
Figure 1: Comparison between previous voxel-based generation (a, b) and the proposed SDF-based FDIF (c), which supports complex topologies and surface-aware textures.
Methodology: The Math of "Synthetic Organs"
The FDIF pipeline follows a rigorous generative process:
- SDF Library: A collection of 109 base classes (spheres, cones, and 106 custom shapes like "hollowed stars").
- Perturbation: Applying Pseudo-Perlin noise or "Sharpmax" bumps to the SDF.
- Intensity Mapping: Using functions like "Inverse Cube" or "Sinusoidal" to fill the 3D volume with intensity.
- Composition: Randomly placing $K$ primitives into a 3D grid and resolving overlaps to create a final "medical-like" volume.
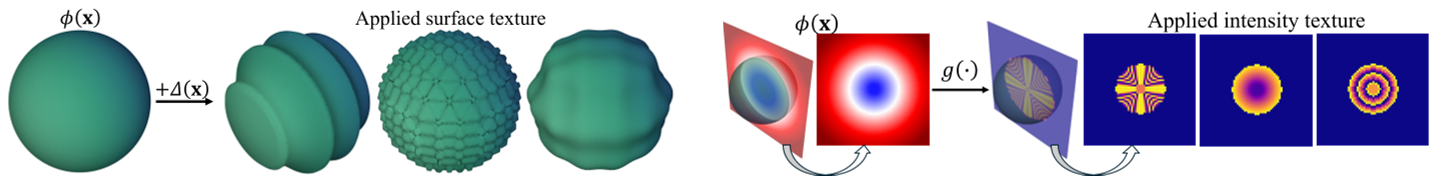 Figure 2: The generation of diverse intensity patterns (Mapper Functions) and geometric surface textures (Displacement Functions).
Figure 2: The generation of diverse intensity patterns (Mapper Functions) and geometric surface textures (Displacement Functions).
Experiments: Beating the Real with the Synthetic
The researchers tested FDIF across three major benchmarks: AMOS (Abdominal), ACDC (Cardiac), and KiTS (Kidney).
Key Findings:
- SOTA in FDSL: Consistently outperformed PrimGeoSeg by +2.58 points on the nnUNet Primus-M architecture.
- Rivaling SSL: On the nnUNet ResEnc-L model, FDIF achieved an average Dice score of 88.70, surpassing MAE (88.30) even though MAE was pre-trained on massive real-world data.
- Generalization: Interestingly, FDIF pre-training also improved performance on 3D Classification tasks (MedMNIST), proving the learned 3D features are robust.
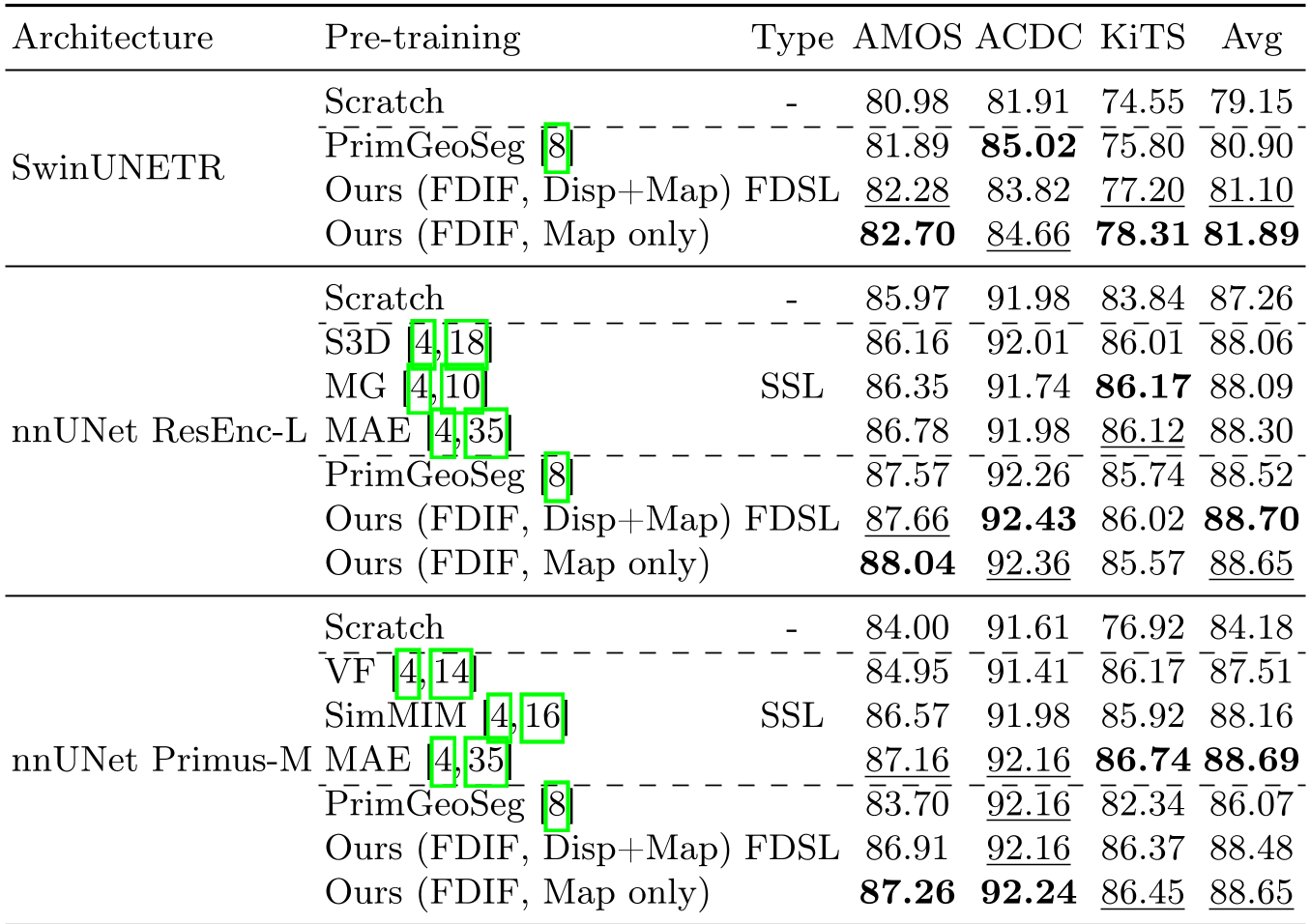 Table 1: Quantative performance across multiple architectures. FDIF (Ours) shows superior or comparable performance to SSL models.
Table 1: Quantative performance across multiple architectures. FDIF (Ours) shows superior or comparable performance to SSL models.
Critical Analysis: Why Does it Work?
The authors' ablation study reveals a profound insight: Texture diversity matters more than shape count. Increasing the number of global shapes (from 10 to 109) only gave a 0.01 gain. However, adding Displacement and Mapper functions provided significantly larger gains.
The Takeaway: Models don't just learn "what an organ looks like"—they learn the underlying principles of 3D geometry and intensity variation.
Conclusion: A Scalable Future
FDIF marks a significant shift in medical AI. By moving away from the "more data" obsession and toward "better math," the researchers have provided a path toward high-performance medical AI that is fundamentally private, scalable, and open.
Limitations: While FDIF is powerful, it still occupies a relatively simple geometric space. Future work could integrate more biologically inspired fractals or even learn the "formulas" themselves from limited real-world samples.
You can access the official code at https://github.com/yamanoko/FDIF.
