The paper introduces MEDOPENCLAW, an auditable runtime that connects Vision-Language Models (VLMs) to medical viewers like 3D Slicer, and MEDFLOW-BENCH, a benchmark for evaluating agents on full 3D clinical studies. It enables models to act as "clinical agents" that navigate multi-sequence MRI/CT/PET scans to reach a diagnosis.
TL;DR
Researchers from TUM and Imperial College London have released MEDOPENCLAW, a runtime that allows AI models to "drive" medical software like 3D Slicer. Alongside it, they introduced MEDFLOW-BENCH, a benchmark that tests if models can navigate full 3D MRI and CT studies. The shock result? Even top-tier models like GPT-4o struggle to use professional tools effectively because they lack the "spatial precision" to click the right pixels.
Background: The "Curation Gap" in Medical AI
Most medical AI today is "cheating." In a typical benchmark, a human radiologist has already found the tumor, cropped the image to a 2D slice, and asked the model: "What is this?" This is Static VQA.
In the real world, a radiologist starts with a "full study"—hundreds of slices across T1, T2, and FLAIR sequences. They must scroll, zoom, and compare. MEDOPENCLAW changes the game by treating the VLM as an Agent that must explore the data itself, creating an auditable trail of evidence.
Methodology: The Auditable Runtime
The core of this work is not a new model, but a Runtime Layer. It acts as a bridge between a Large Vision-Language Model (LVLM) and 3D Slicer.
1. Three-Layer Action Space
- Primitive Actions: Scrolling, selecting sequences, changing window levels (brightness/contrast).
- Evidence Operations: Bookmarking specific slices or drawing masks.
- Expert Tools: Invoking advanced algorithms (e.g., MONAI segmentation).
2. Guardrailed Auditability
Unlike agents that generate raw Python code (which is risky and hard to audit), MEDOPENCLAW uses a bounded API. The model sends a command (e.g., scroll_to_slice(42)), and the system logs it. This creates a "replayable" diagnostic trajectory that a human doctor can review.
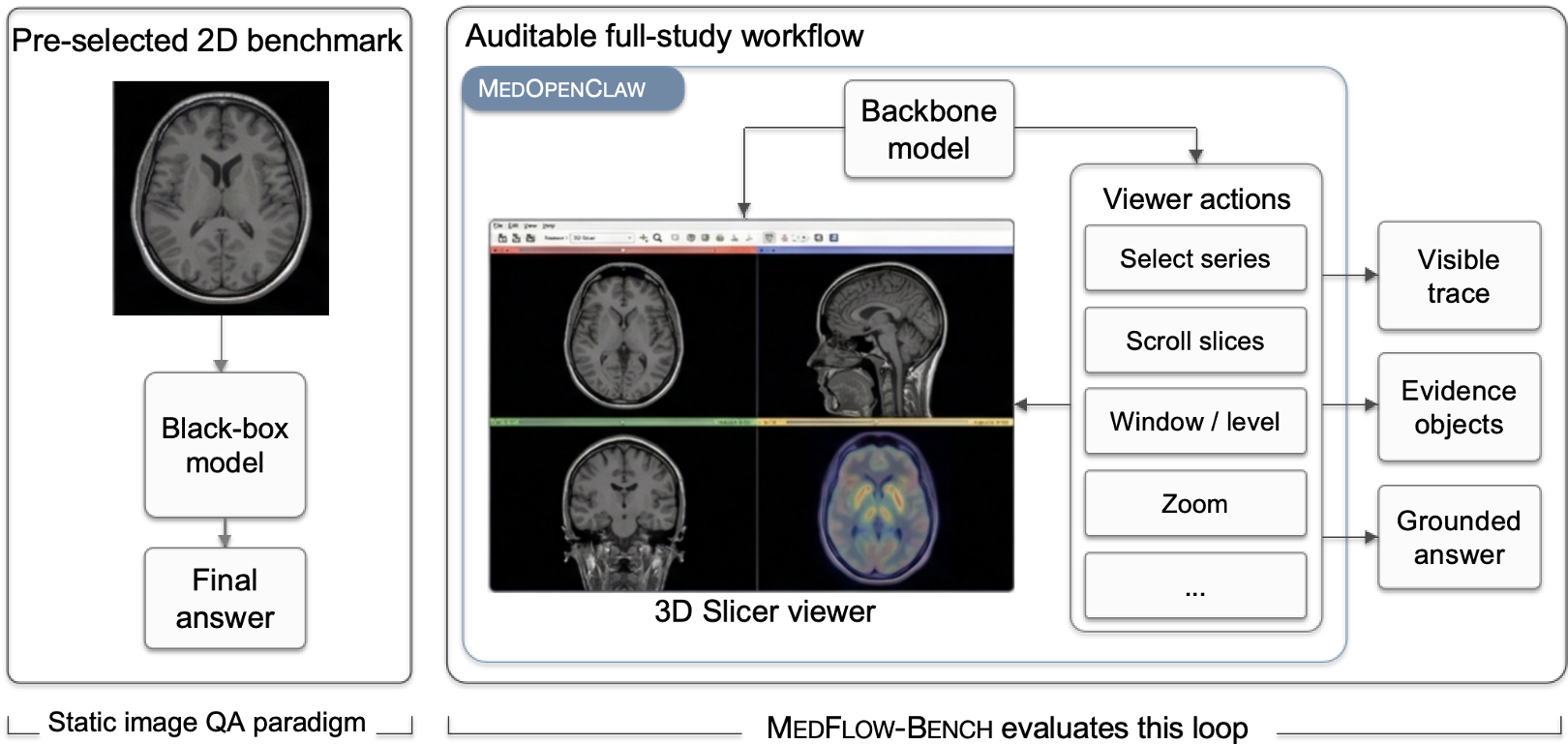 The shift from black-box static VQA (left) to the auditable, interactive loop of MEDOPENCLAW (right).
The shift from black-box static VQA (left) to the auditable, interactive loop of MEDOPENCLAW (right).
The Benchmark: MEDFLOW-BENCH
The authors tested the world's leading models on two complex tasks:
- Brain MRI (UCSF-PDGM): Diagnosing tumor types from multi-sequence scans.
- Lung CT/PET (NSCLC): Detailed cancer staging and pathology.
Results: The Tool-Use Paradox
The results revealed a fascinating and counter-intuitive trend.
| Backbone Model | Viewer-Only Accuracy | With Segmentation Tools | | :--- | :---: | :---: | | GPT-4o (5.4) | 0.32 | 0.27 (↓) | | GPT-4o-mini | 0.20 | 0.14 (↓) |
Why did performance drop when the models got "better" tools? The researchers call this the spatial grounding bottleneck. To use a segmentation tool, a model must provide exact 3D coordinates. If the model is off by even a few millimeters, the tool segmentates the wrong tissue. The model then looks at its own "fake" evidence and becomes even more confident in a wrong answer.
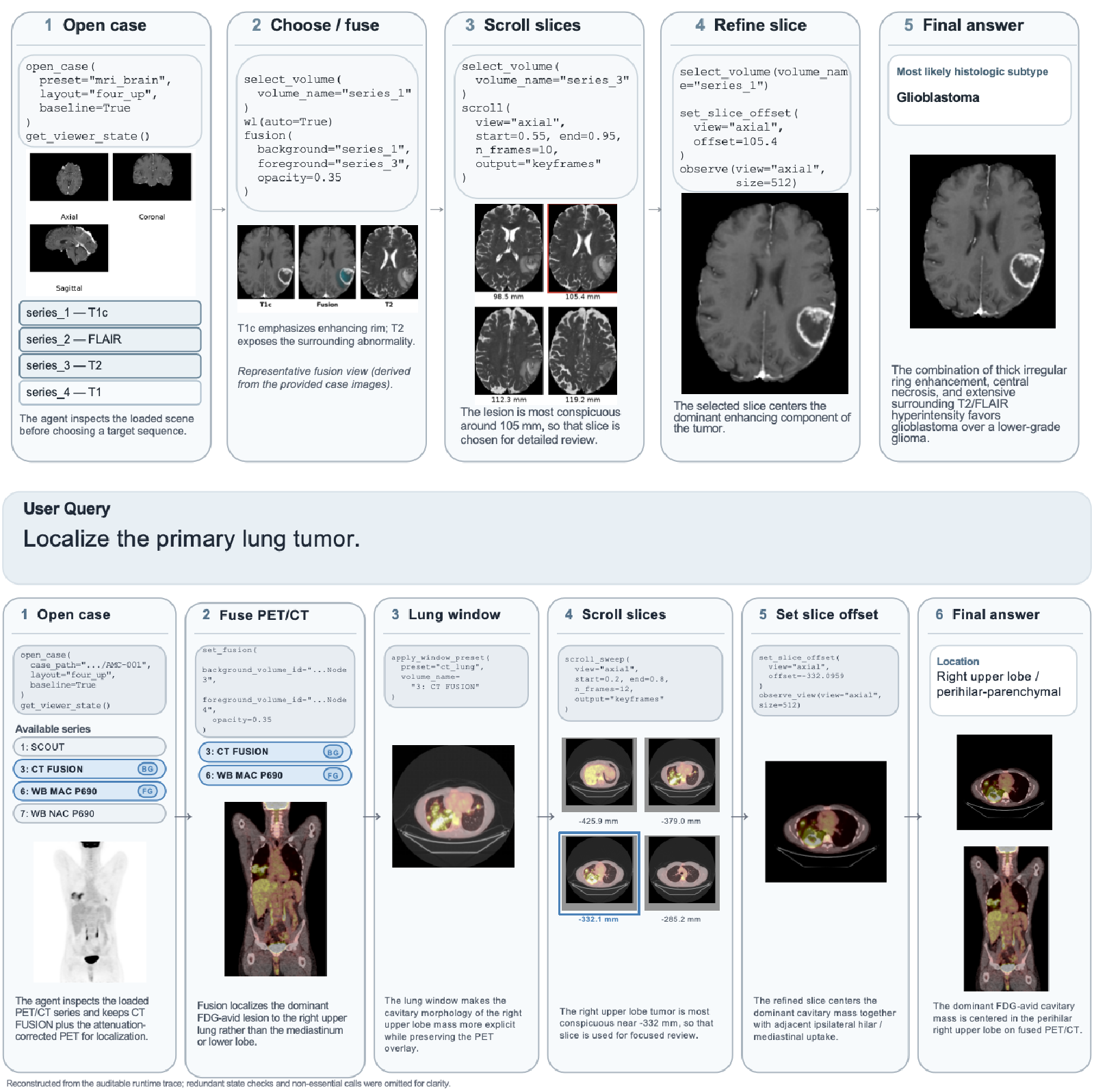 A step-by-step trace showing the agent gathering evidence across different MRI sequences.
A step-by-step trace showing the agent gathering evidence across different MRI sequences.
Critical Insight: The Road to Clinical Trust
The paper makes a powerful argument: Auditability is not a feature; it is a prerequisite.
For AI to enter the hospital, "The model said so" is not enough. We need to see that the model:
- Checked the T2 sequence to look for edema.
- Compared the CT and PET scans to find hypermetabolic activity.
- Measured the tumor using standard tools.
MEDOPENCLAW proves that while SOTA models can navigate (the "Viewer-Only" track), they are not yet "digital surgeons" capable of surgical precision in tool interaction.
Conclusion and Future Work
The authors have laid the foundation for MEDCOPILOT, a system where AI handles the "grunt work" of radiology (fusing images, finding key slices) while the human doctor focuses on the final interpretation.
The Next Frontier: Improving Spatial Grounding. We need vision models that don't just "describe" images but understand the 3D coordinate geometry of the human body.
Disclaimer: This analysis is based on the MEDOPENCLAW paper (2024). All images and data credits belong to the original authors.
