VolDiT is the first purely transformer-based 3D Diffusion Transformer (DiT) specifically designed for volumetric medical image synthesis. By replacing traditional convolutional U-Net backbones with global self-attention over 3D latent tokens, it achieves state-of-the-art performance in generating high-fidelity Lung CT and Cardiac CTA volumes.
TL;DR
VolDiT is a pioneering framework that replaces the standard 3D U-Net backbone in diffusion models with a pure 3D Transformer architecture. It leverages global self-attention to model volumetric data (like CT scans) and introduces a Timestep-Gated Control Adapter (TGCA) for precise, mask-guided generation. It represents a significant shift toward scalable, token-based architectures in medical AI.
The Problem: The Locality Trap of 3D U-Nets
For years, the 3D U-Net has been the workhorse of medical imaging. However, its reliance on local convolutions creates a "locality bias." In volumetric imaging, where anatomical structures (like the aorta or lungs) span across many slices, a global understanding is crucial. Traditional U-Nets often struggle with:
- Limited Receptive Fields: Difficulty in maintaining long-range anatomical consistency.
- Scale-Up Bottlenecks: Convolutional architectures don't always translate increased parameter counts into better sample quality as efficiently as Transformers.
- Rigid Conditioning: Integrating complex spatial priors (like segmentation masks) often requires heavy architectural modifications.
Methodology: The 3D DiT Edge
VolDiT addresses these issues by treating a 3D volume not as a stack of images, but as a sequence of 3D tokens.
1. Volumetric Tokenization
The process begins with a 3D VQ-GAN that compresses the medical volume into a latent space. VolDiT then divides this latent space into non-overlapping cubic patches (e.g., $2 imes 2 imes 2$ or $4 imes 4 imes 4$). These patches are flattened and embedded, allowing the model to apply Global Self-Attention across the entire volume simultaneously.
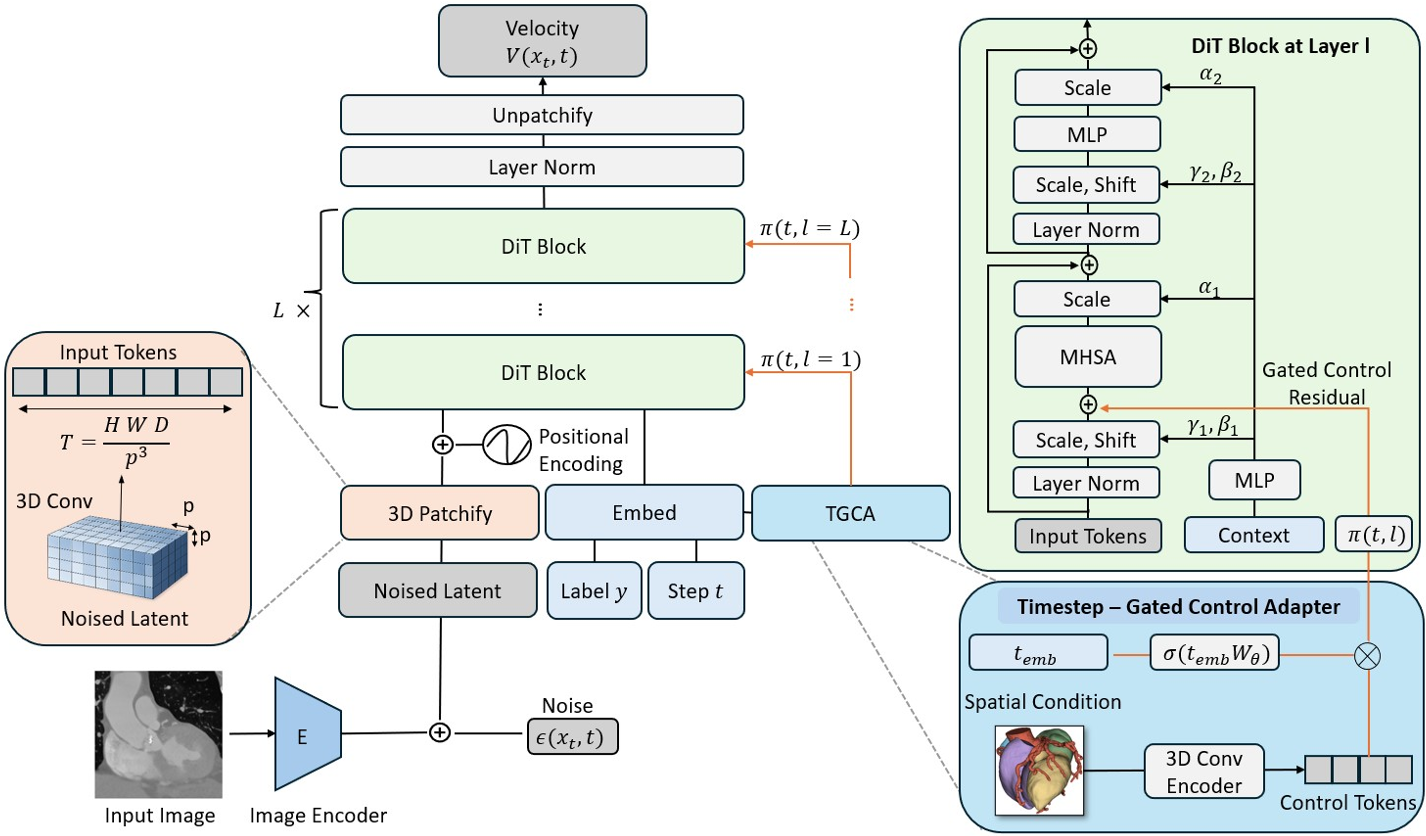
2. Timestep-Gated Control (TGCA)
To make the synthesis controllable, the authors developed the TGCA. Unlike static adapters, TGCA uses a learnable gating mechanism $\gamma(t)$ that adjusts the influence of the conditioning signal (e.g., a segmentation mask) depending on the diffusion timestep. This allows the model to focus on structure during early denoising stages and texture in the final stages.
Experimental Performance
The researchers tested VolDiT on two challenging datasets: LUNA16 (Lung CT) and TaviCT (Cardiac CTA).
Performance vs. Baselines
Compared to state-of-the-art U-Net LDMs and HA-GANs, VolDiT showed a clear advantage in both fidelity (FID) and diversity (MS-SSIM).
| Dataset | Model | FID (↓) | Precision (↑) | Recall (↑) | | :--- | :--- | :--- | :--- | :--- | | TaviCT | U-Net LDM | 36.8 | 0.95 | 0.63 | | TaviCT | VolDiT (Ours) | 21.4 | 0.94 | 0.73 |
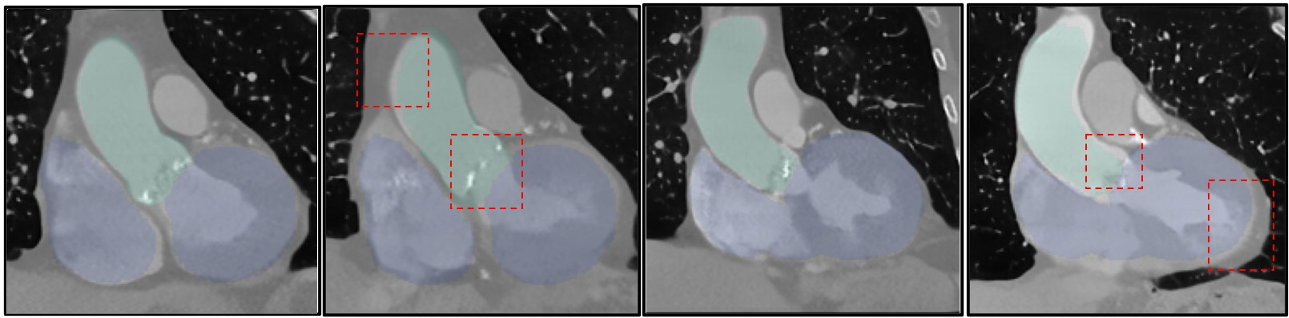 Fig 3: Samples generated by VolDiT show much tighter alignment with anatomical masks compared to the U-Net baseline, which often lacks structural realism in constrained areas.
Fig 3: Samples generated by VolDiT show much tighter alignment with anatomical masks compared to the U-Net baseline, which often lacks structural realism in constrained areas.
The Scaling Paradox
Interestingly, the authors noted that while DiT models are known to scale perfectly in 2D Natural Images, the improvement was less "monotonic" in 3D medical data. This highlights a critical insight: Medical datasets are often smaller than ImageNet, and large-scale Transformers might require longer training or better initialization to reach their full potential in this domain.
Critical Insight & Future Outlook
The most profound takeaway from VolDiT is the architectural unification. By moving to a token-based backbone, medical imaging can finally "speak the same language" as modern LLMs and Multimodal Foundation Models.
Limitations:
- The computational cost of global self-attention on very high-resolution 3D volumes is still high.
- Sensitivity to the autoencoder's compression rate.
The Future: VolDiT sets the stage for a 4D extension (spatiotemporal cardiac modeling) and suggests that the future of medical synthesis lies in flexible, transformer-based architectures that can ingest any modality or anatomical prior through a simple token interface.
